Recombinant Human DNA mismatch repair protein Mlh1 (MLH1)
-
货号:CSB-YP014624HU
-
规格:
-
来源:Yeast
-
其他:
-
货号:CSB-EP014624HU
-
规格:
-
来源:E.coli
-
其他:
-
货号:CSB-EP014624HU-B
-
规格:
-
来源:E.coli
-
共轭:Avi-tag Biotinylated
E. coli biotin ligase (BirA) is highly specific in covalently attaching biotin to the 15 amino acid AviTag peptide. This recombinant protein was biotinylated in vivo by AviTag-BirA technology, which method is BriA catalyzes amide linkage between the biotin and the specific lysine of the AviTag.
-
其他:
-
货号:CSB-BP014624HU
-
规格:
-
来源:Baculovirus
-
其他:
-
货号:CSB-MP014624HU
-
规格:
-
来源:Mammalian cell
-
其他:
产品详情
-
纯度:>85% (SDS-PAGE)
-
基因名:
-
Uniprot No.:
-
别名:COCA 2; COCA2; DNA mismatch repair protein Mlh1; FCC 2; FCC2; hMLH 1; hMLH1; HNPCC 2; HNPCC; HNPCC2; MGC5172; MLH 1; MLH1; MLH1_HUMAN; MutL homolog 1 (E. coli); MutL homolog 1; MutL homolog 1 colon cancer nonpolyposis type 2; MutL homolog 1, colon cancer, nonpolyposis type 2 (E. coli); MutL protein homolog 1; MutL, E. coli, homolog of, 1
-
种属:Homo sapiens (Human)
-
蛋白长度:Full length protein
-
表达区域:1-756
-
氨基酸序列MSFVAGVIRR LDETVVNRIA AGEVIQRPAN AIKEMIENCL DAKSTSIQVI VKEGGLKLIQ IQDNGTGIRK EDLDIVCERF TTSKLQSFED LASISTYGFR GEALASISHV AHVTITTKTA DGKCAYRASY SDGKLKAPPK PCAGNQGTQI TVEDLFYNIA TRRKALKNPS EEYGKILEVV GRYSVHNAGI SFSVKKQGET VADVRTLPNA STVDNIRSIF GNAVSRELIE IGCEDKTLAF KMNGYISNAN YSVKKCIFLL FINHRLVEST SLRKAIETVY AAYLPKNTHP FLYLSLEISP QNVDVNVHPT KHEVHFLHEE SILERVQQHI ESKLLGSNSS RMYFTQTLLP GLAGPSGEMV KSTTSLTSSS TSGSSDKVYA HQMVRTDSRE QKLDAFLQPL SKPLSSQPQA IVTEDKTDIS SGRARQQDEE MLELPAPAEV AAKNQSLEGD TTKGTSEMSE KRGPTSSNPR KRHREDSDVE MVEDDSRKEM TAACTPRRRI INLTSVLSLQ EEINEQGHEV LREMLHNHSF VGCVNPQWAL AQHQTKLYLL NTTKLSEELF YQILIYDFAN FGVLRLSEPA PLFDLAMLAL DSPESGWTEE DGPKEGLAEY IVEFLKKKAE MLADYFSLEI DEEGNLIGLP LLIDNYVPPL EGLPIFILRL ATEVNWDEEK ECFESLSKEC AMFYSIRKQY ISEESTLSGQ QSEVPGSIPN SWKWTVEHIV YKALRSHILP PKHFTEDGNI LQLANLPDLY KVFERC
-
蛋白标签:Tag type will be determined during the manufacturing process.
The tag type will be determined during production process. If you have specified tag type, please tell us and we will develop the specified tag preferentially. -
产品提供形式:Lyophilized powder
Note: We will preferentially ship the format that we have in stock, however, if you have any special requirement for the format, please remark your requirement when placing the order, we will prepare according to your demand. -
复溶:We recommend that this vial be briefly centrifuged prior to opening to bring the contents to the bottom. Please reconstitute protein in deionized sterile water to a concentration of 0.1-1.0 mg/mL.We recommend to add 5-50% of glycerol (final concentration) and aliquot for long-term storage at -20℃/-80℃. Our default final concentration of glycerol is 50%. Customers could use it as reference.
-
储存条件:Store at -20°C/-80°C upon receipt, aliquoting is necessary for mutiple use. Avoid repeated freeze-thaw cycles.
-
保质期:The shelf life is related to many factors, storage state, buffer ingredients, storage temperature and the stability of the protein itself.
Generally, the shelf life of liquid form is 6 months at -20°C/-80°C. The shelf life of lyophilized form is 12 months at -20°C/-80°C. -
货期:Delivery time may differ from different purchasing way or location, please kindly consult your local distributors for specific delivery time.Note: All of our proteins are default shipped with normal blue ice packs, if you request to ship with dry ice, please communicate with us in advance and extra fees will be charged.
-
注意事项:Repeated freezing and thawing is not recommended. Store working aliquots at 4°C for up to one week.
-
Datasheet :Please contact us to get it.
相关产品
靶点详情
-
功能:Heterodimerizes with PMS2 to form MutL alpha, a component of the post-replicative DNA mismatch repair system (MMR). DNA repair is initiated by MutS alpha (MSH2-MSH6) or MutS beta (MSH2-MSH3) binding to a dsDNA mismatch, then MutL alpha is recruited to the heteroduplex. Assembly of the MutL-MutS-heteroduplex ternary complex in presence of RFC and PCNA is sufficient to activate endonuclease activity of PMS2. It introduces single-strand breaks near the mismatch and thus generates new entry points for the exonuclease EXO1 to degrade the strand containing the mismatch. DNA methylation would prevent cleavage and therefore assure that only the newly mutated DNA strand is going to be corrected. MutL alpha (MLH1-PMS2) interacts physically with the clamp loader subunits of DNA polymerase III, suggesting that it may play a role to recruit the DNA polymerase III to the site of the MMR. Also implicated in DNA damage signaling, a process which induces cell cycle arrest and can lead to apoptosis in case of major DNA damages. Heterodimerizes with MLH3 to form MutL gamma which plays a role in meiosis.
-
基因功能参考文献:
- Low MLH1 expression is associated with Colonic Adenoma and Adenocarcinoma. PMID: 29976631
- Single-Nucleotide Polymorphisms of the MLH1 is associated with Basal Cell Carcinoma. PMID: 28667494
- this meta-analysis suggested that hMLH1 methylation was associated with an increased CRC risk. PMID: 29970664
- The DNA mismatch repair gene (MMR) plays an important role in anti-carcinogenesis and MMR gene deficiency. PMID: 29728922
- Study investigated the counteraction of oxidative stress by vitamin E in the colorectal cancer cell line Caco-2 under normal and high glucose cell culture condition. Gene expression and promoter methylation of the DNA repair gene MutL homolog 1 (MLH1) and the DNA methyltransferase 1 (DNMT1) were investigated. Induction of MLH1 and DNMT1 gene expression was noticed, accompanied by an increase in global methylation. PMID: 29854080
- the results identified for the first time a MLH1 missense mutation (NM_000249.3:p.Tyr379Ser/c.1136A>C) in a Chinese family with Gardner syndrome (GS), thus broadening the range of mutated genes associated with GS. PMID: 29845239
- This study demonstrated that Aberrant methylation of mutL homolog 1 is associated with increased risk of non-small cell lung cancer PMID: 29205508
- MLH1's ATPase domain has a crucial role in preventing the aberrant formation of telomeric sequences at the intra-chromosomal regions and preserving genome stability. PMID: 29544212
- provide evidence that four of the alterations in MLH1 are causative for Lynch syndrome, four are likely neutral and one shows compromised activity which can currently not be classified with respect to its pathogenic potential PMID: 29520894
- This pilot study indicates oligozoospermic patients have more methylation of MLH1 than normozoospermic patients. PMID: 28983945
- MLH1 and PMS2 can be imported to the nucleus by a classical nuclear import pathway PMID: 29175432
- The MLH1-93 AA genotype is significantly associated with promoter hypermethylation and MLH1 loss in the context of Sessile serrated adenoma of dysplasia. BRAF mutant microsatellite stable colorectal cancers with the AA genotype most likely arise in traditional serrated adenomas since the A allele does not predispose to methylation in this context. PMID: 29304767
- Cancer developed more often in mutation carriers, with no consistent difference between MLH1 and MSH2 carriers. More polyps (mostly adenomas) were detected in MLH1 carriers. The majority (13 of 21) of malignant tumours occurred in organs for which there is no recommended surveillance, and were lethal in three patients. PMID: 29025352
- Data suggest that the concurrent mutations of the adenomatous polyposis coli protein (APC) and mutL protein homolog 1 (MLH1) genes probably underline the familial adenomatous polyposis (FAP) in the pedigree. PMID: 29419868
- Among 13 gastric tumors showing no hMLH1 or hMSH2 expression, 8 MSI-H (high) and 5 MSI-L (low) were identified. PMID: 28525661
- Results show that folate concentrations >45.3 nmol/L increased the risk of methylation in specific CpG sites of MLH1. PMID: 28748002
- In contrast to colorectal carcinoma, sporadic MLH1 deficiency is not seen in small bowel adenocarcinoma PMID: 27258561
- MLH1 methylation is associated with colorectal cancer. PMID: 28730335
- We found no correlation between the DLEC1, TUSC4 and MLH1 gene expression and NSCLC patient characteristics (gender, age and smoking) or cancer histopathology. No significant differences in the gene expression among NSCLC subtypes indicate the weakness of DLEC1, TUSC4 and MLH1 expression analysis as potential differentiating markers of NSCLC subtypes in the Polish population. PMID: 28674222
- The results of this meta-analysis show a significant association between hMLH1 hypermethylation and colorectal cancer risk. PMID: 28635682
- study to estimate frequency of point mutations and chromosomal rearrangements in 3 mismatch repair (MMR) genes MLH1, MSH2, and MSH6 among unselected patients with ovarian cancer; estimate that approximately 0.6% of unselected ovarian cancer patients have mutations in the MMR genes PMID: 28176205
- CpG island methylator phenotype (CIMP)+/MLH1-unmethylated (MLH1-U) tumors exhibit aggressive features and are associated with poor clinical outcomes. PMID: 27880934
- Analysis of expression Quantitative Trait Loci (eQTLs) provides modest support for altered regulation of MLH1 and LRRFIP2, raising the possibility that the modifier affects regulation of both genes. PMID: 28934397
- MLH1 deficiency had an impact on the evolution of the tumor after surgical resection in a murine xenograft model. PMID: 28512763
- Results indicate that the normal tissue types tested (colorectum and PBMC) experience dynamic genotype-associated epigenetic alterations at the MLH1 shore, whereas cancer tumor DNA incurs aberrant hypermethylation compared to normal DNA. These results reveal that the epigenetic landscape of MLH1 is dynamically regulated at least in part by the static genetic sequence. PMID: 28293327
- MLH1-methylated CpG island methylator phenotype is associated with serrated adenoma in colorectal carcinoma. PMID: 26883113
- cancer-specific survival did not differ in Lynch syndrome-associated colorectal cancer (CRC) compared with MLH1-hm CRC, suggesting that they carry a similar prognosis. PMID: 26866578
- Data show that MLH1(L749P) and MLH1(Y750X) make PMS2 prone to calyculin induced degradation, suggeting that the specific degradation of PMS2 may represent a new mechanism to regulate MLH1-PMS2 heterodimer protein MutLalpha. PMID: 28767177
- hMLH1 (rs1800734) AA genotype is associated with increased risk of lung cancer development. PMID: 26987333
- miR-148b mimic sensitized A549 xenografts to irradiation in vivo Our study demonstrated that miR-148b regulates radioresistance of lung cancer cells by modulating MLH1 expression level. miR-148b may represent a new therapeutic target for the intervention of lung cancer. PMID: 26759383
- hMLH1 promoter hypermethylation plays important roles in esophageal carcinoma drug-resistance PMID: 26782688
- Loss of MLH1 expression is associated with colorectal carcinoma. PMID: 28651545
- the MLH1 c.543C>T silent mutation is considered 'pathogenic'. PMID: 28334867
- Promoter methylation and loss of protein expression of hMLH1 are not parallel processes that occur concurrently. hMLH1 methylation is an early molecular event which occurs even in hyperplastic polyps (HPs). the loss of hMLH1 protein expression does not necessarily precede the development of cytological dysplasia in sessile serrated lesion (SSL). PMID: 26713412
- Overexpression of MutL homolog 1 and MutS homolog 2 proteins have reversed prognostic implications for stage I-II colon cancer PMID: 28411881
- MLH1, a key protein involved in mismatch repair (MMR), suppresses telomeric sequence insertion (TSI) at intra-chromosomal regions. The frequency of TSI can be elevated by double-strand break (DSB) inducer and abolished by ATM/ATR inhibition. Suppression of TSI requires MLH1 recruitment to DSBs, indicating that MLH1's role in DSB response/repair is important for suppressing TSI. PMID: 28180301
- MLH1 mutations contribute to colorectal cancer susceptibility in Algerian families with suspected Lynch syndrome. PMID: 27468915
- MLH1 methylation analysis defines a subset of tumors that have worse prognostic features including higher tumor volume and lymph node involvement. PMID: 28709704
- High MLH1 expression is associated with the development of genetic instability and is linked to tumor aggressiveness and early PSA recurrence in prostate cancer. PMID: 27803051
- In path_MLHI 10-year survival was 80%, relative incidence of subsequent cancer compared with incidence of first cancer was slightly but insignificantly higher than cancer incidence in patients with Lynch syndrome without previous cancer. PMID: 27261338
- MLH1 methylation is a frequent molecular event in colorectal cancer and lung cancer patients PMID: 28214209
- Both the mTOR/p-mTOR and BNIP3/Beclin-1 signaling pathways were found to be related to HRS, but only mTOR/p-mTOR is involved in the regulation of HRS via MLH1 and autophagy. PMID: 28117625
- Data suggest that yeast Mlh1-Mlh3 heterodimer does not exhibit hallmarks of a canonical (structure-selective) Holliday junction resolvase/endonuclease; multiple Mlh1-Mlh3 heterodimers appear to load onto DNA to form an activated polymer that cleaves DNA; human MLH1-PMS2 exhibits similar characteristics. (MLH = mutL homolog protein; PMS2 = post meiotic segregation increased 2 protein) PMID: 28453523
- Genetic changes between MLH1 and MSH2 were significantly positively correlated (p = 0.032). We also noted a positive correlation between genetic changes of MSH2 and DVL3 genes (p = 0.034). PMID: 28705114
- Mismatch repair (MMR)-deficient endometrial carcinomas have significantly increased tumoral and immune stromal PD-L1 expression relative to MMR-intact carcinomas. PD-L1 is frequently positive in both tumoral and immune cells at the periphery of MMR-deficient tumors. Lynch syndrome-associated tumors are more likely to express PDL1 relative to sporadic MLH1 promoter hypermethylation carcinomas. PMID: 27984238
- These results demonstrate the role of variant rs1800734 in altering transcription factor binding as well as epigenetics at regions beyond the MLH1 CpG island in which it is located. PMID: 28304185
- There were a 61.7% (95/154) overexpression of DNMT3B, 50.0% (77/154) loss of PTEN expression and 18.2% (28/154) loss of hMLH1 expression in endometrial carcinomas. PMID: 28220037
- demonstrates that lower level of histone acetylation and specific histone methylation such as H3K36me3 correlate with aberrant transcripts in hMLH1 exon 10-11 region PMID: 28381181
- revealed high frequencies of gene promoter methylation in NSCLCs: 84 % for DLEC1 and MLH1 and 57 % for ITGA9 PMID: 27287342
- In colorectal neoplasms, negative expression of the MMR proteins MLH1, MSH2 or MSH6 was seen in 15% (47 of 313) of the patients. Defect MLH1 was most common and detected in 12% of the cases. Defect MLH1 and MSH2 were identified in each patient's normal adjacent mucosa. PMID: 27836416
显示更多
收起更多
-
相关疾病:Hereditary non-polyposis colorectal cancer 2 (HNPCC2); Mismatch repair cancer syndrome (MMRCS); Muir-Torre syndrome (MRTES); Endometrial cancer (ENDMC); Colorectal cancer (CRC)
-
亚细胞定位:Nucleus. Chromosome.
-
蛋白家族:DNA mismatch repair MutL/HexB family
-
组织特异性:Colon, lymphocytes, breast, lung, spleen, testis, prostate, thyroid, gall bladder and heart.
-
数据库链接:
HGNC: 7127
OMIM: 114500
KEGG: hsa:4292
STRING: 9606.ENSP00000231790
UniGene: Hs.195364
Most popular with customers
-
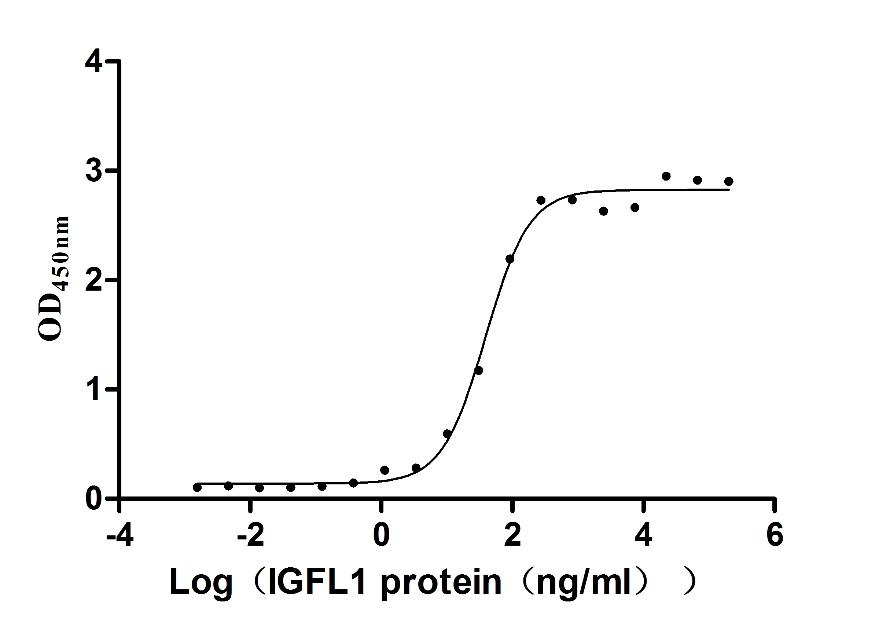
Recombinant Human Insulin growth factor-like family member 1 (IGFL1) (Active)
Express system: Mammalian cell
Species: Homo sapiens (Human)
-
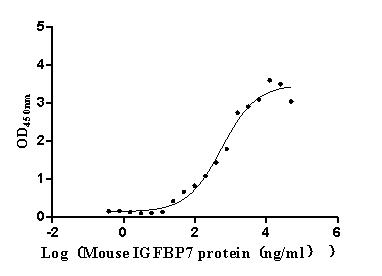
Recombinant Mouse Complement component C1q receptor (Cd93), partial (Active)
Express system: Mammalian cell
Species: Mus musculus (Mouse)
-
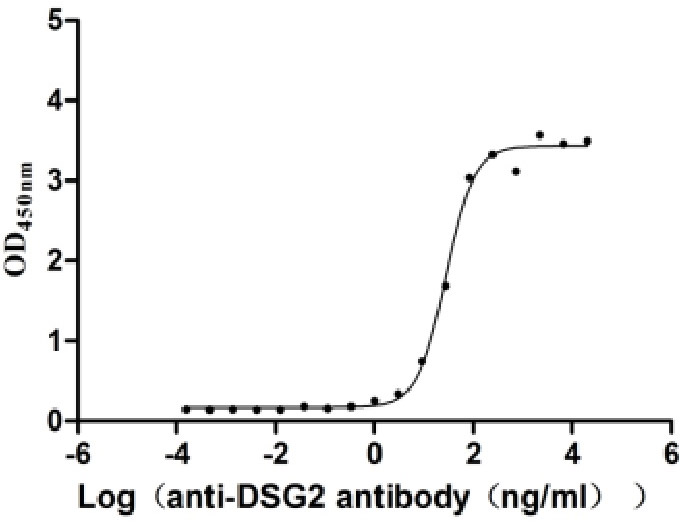
Recombinant Human Desmoglein-2 (DSG2), partial (Active)
Express system: Mammalian cell
Species: Homo sapiens (Human)
-
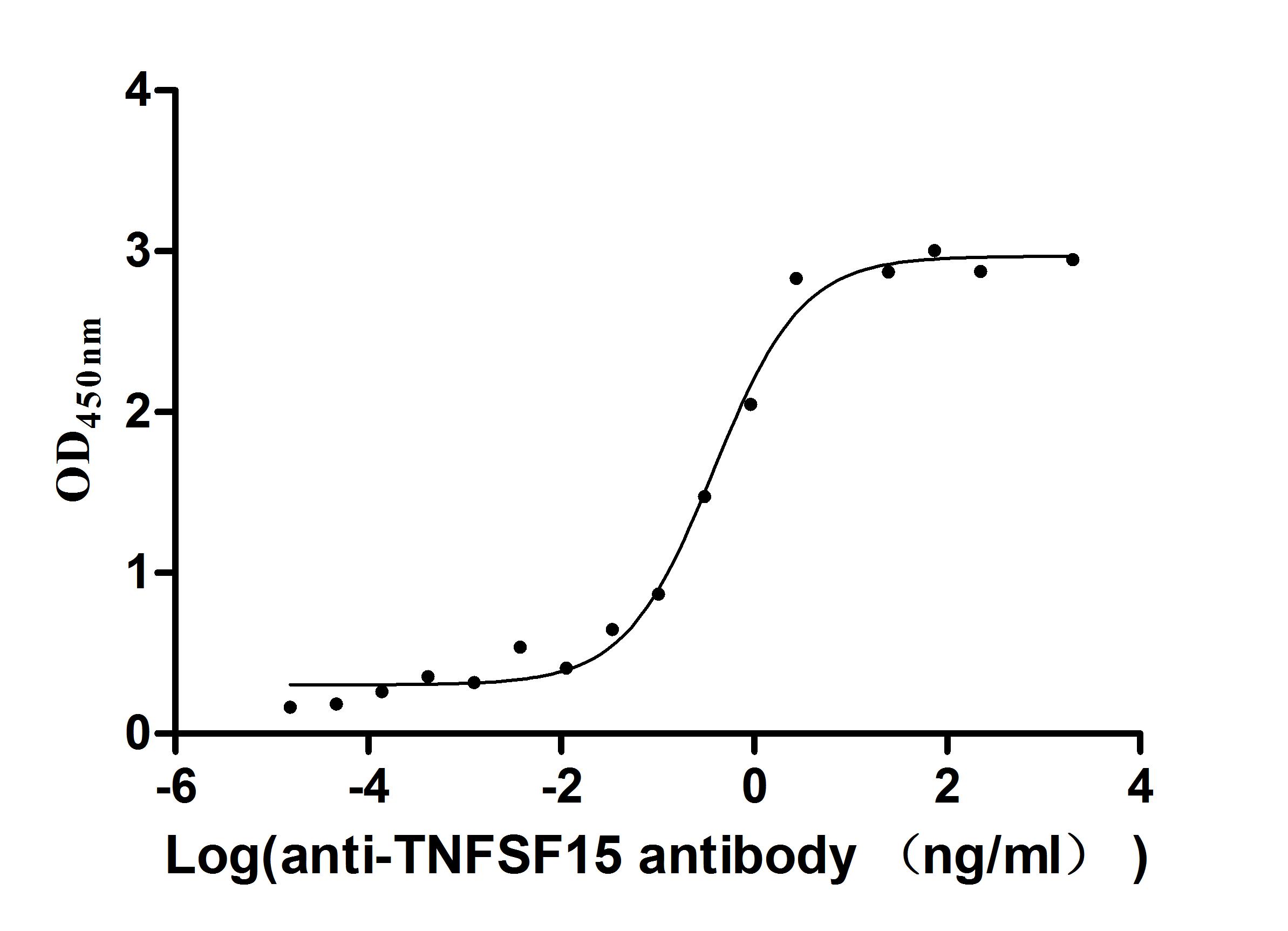
Express system: Mammalian cell
Species: Homo sapiens (Human)
-
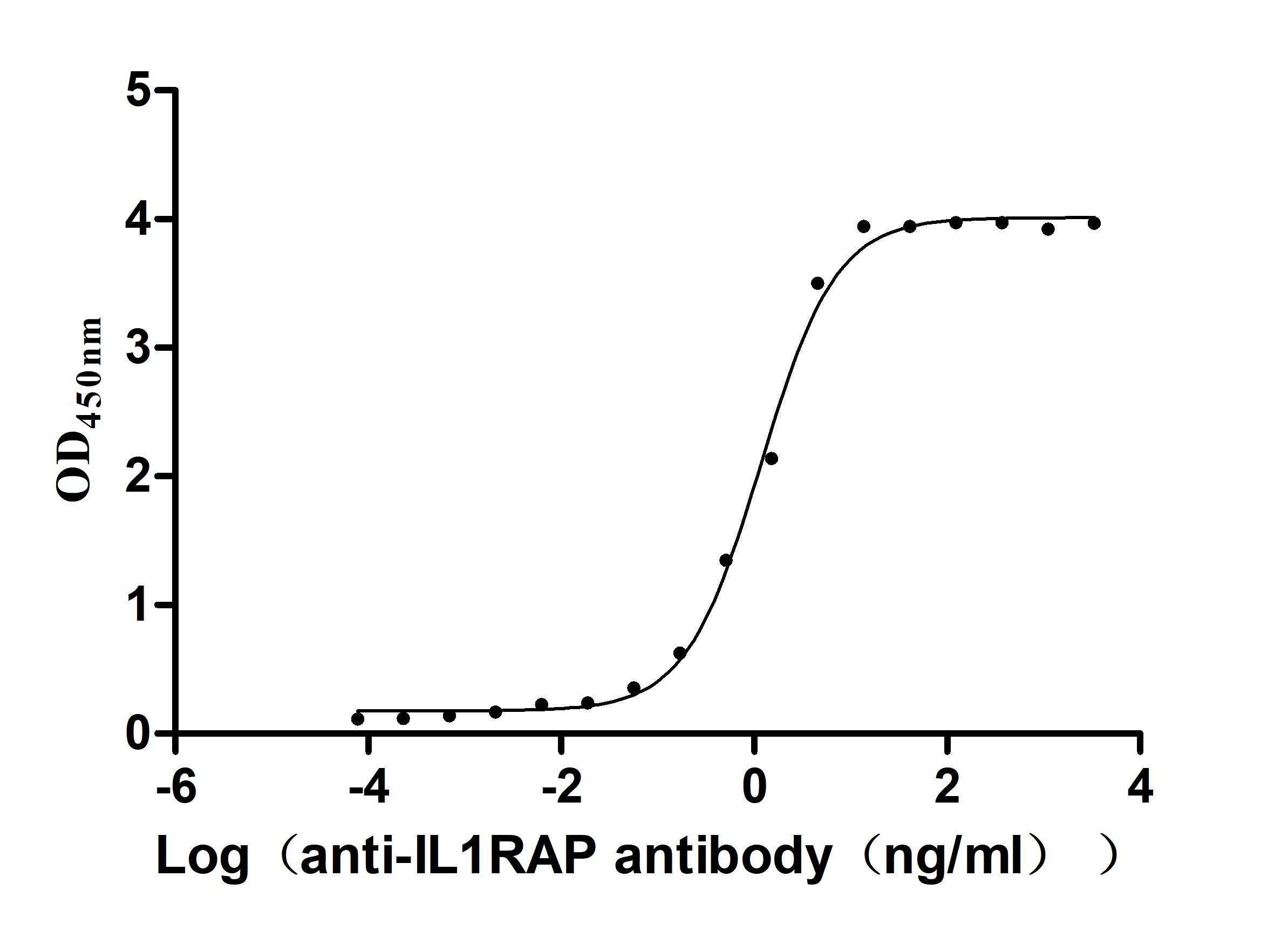
Recombinant Human Interleukin-1 receptor accessory protein (IL1RAP), partial (Active)
Express system: Mammalian cell
Species: Homo sapiens (Human)